Note
Go to the end to download the full example code.
Introduction to using queues
This demonstrates how to use queues to order stimuli
import pylab as plt
import numpy as np
from psiaudio import stim
from psiaudio.calibration import FlatCalibration
from psiaudio.queue import FIFOSignalQueue
from psiaudio.queue import (BlockedRandomSignalQueue, FIFOSignalQueue,
InterleavedFIFOSignalQueue)
fs = 20000
First, let’s create factories for our stimuli (chirp, bandlimited noise bursts, and tones).
calibration = FlatCalibration.as_attenuation()
chirp = stim.ChirpFactory(fs, start_frequency=5, end_frequency=50, duration=1,
level=0, calibration=calibration)
noise = stim.BandlimitedNoiseFactory(fs, seed=0, level=0, fl=100, fh=1000,
filter_rolloff=2, passband_attenuation=1,
stopband_attenuation=80,
calibration=calibration)
noise_burst = stim.Cos2EnvelopeFactory(fs, duration=1, rise_time=0.25,
input_factory=noise)
tone = stim.ToneFactory(fs, frequency=3, level=0, calibration=calibration)
tone_burst = stim.Cos2EnvelopeFactory(fs, duration=1, rise_time=0.25,
input_factory=tone)
Now, create a first-in-first-out queue with five trials of each stimulus. This meanst that five chirps will be presented, followed by five noise bursts, followed by five tone bursts. The delays specifies the intertrial interval.
queue = FIFOSignalQueue(fs)
uuid = queue.append(chirp, trials=5, delays=1)
uuid = queue.append(noise_burst, trials=5, delays=1)
uuid = queue.append(tone_burst, trials=5, delays=1)
Whenever you append a new stimulus to the queue, a universally unique identifier (UUID) is returned. You can use that as an internal reference to track what stimuli are generated
uuid
'd23cb901-4f7d-49f1-9687-40f35c2352ff'
Since each stimulus is 1 second long with a 1 second intertrial interval and there are five trials of three stimuli, the entire train will take 30 seconds. Let’s “pop” that off of the queue buffer.
y = queue.pop_buffer(samples=fs*30)
t = np.arange(len(y)) / fs
plt.plot(t, y)
plt.xlabel('Time (s)')
plt.show()
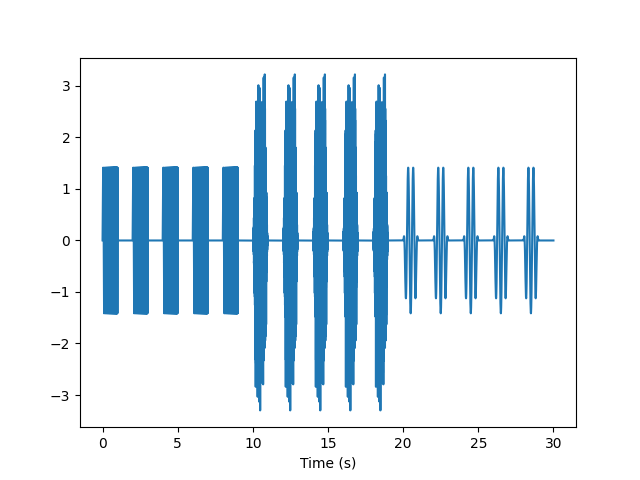
Want to interleave the stimuli? Use InterleavedFIFOSignalQueue.
queue = InterleavedFIFOSignalQueue(fs=fs)
queue.append(chirp, trials=5, delays=1)
queue.append(noise_burst, trials=5, delays=1)
queue.append(tone_burst, trials=5, delays=1)
y = queue.pop_buffer(samples=fs*30)
t = np.arange(len(y)) / fs
plt.plot(t, y)
plt.xlabel('Time (s)')
plt.show()
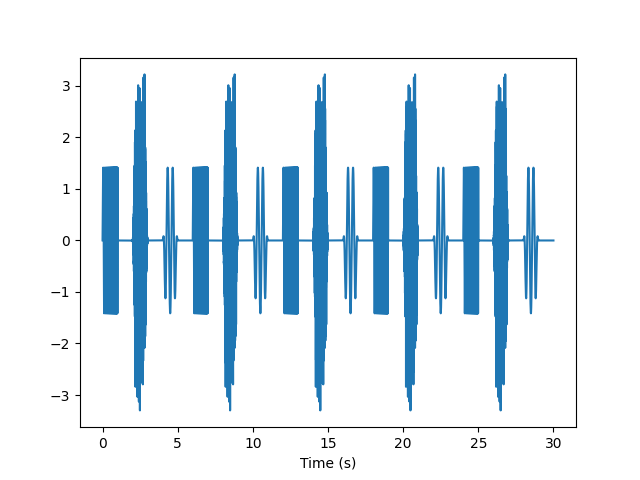
There are many interesting permutations on the queueing mechanism. We can present the set of stimuli in random order, but ensure that in each “block” of three trials all three stimuli are presented (but in random order).
# Set the random seed to 1 since this creates an ordering that is clearly
# semi-randomized.
queue = BlockedRandomSignalQueue(fs=fs, seed=1)
queue.append(chirp, trials=5, delays=1)
queue.append(noise_burst, trials=5, delays=1)
queue.append(tone_burst, trials=5, delays=1)
y = queue.pop_buffer(samples=fs*30)
t = np.arange(len(y)) / fs
plt.plot(t, y)
plt.xlabel('Time (s)')
plt.show()
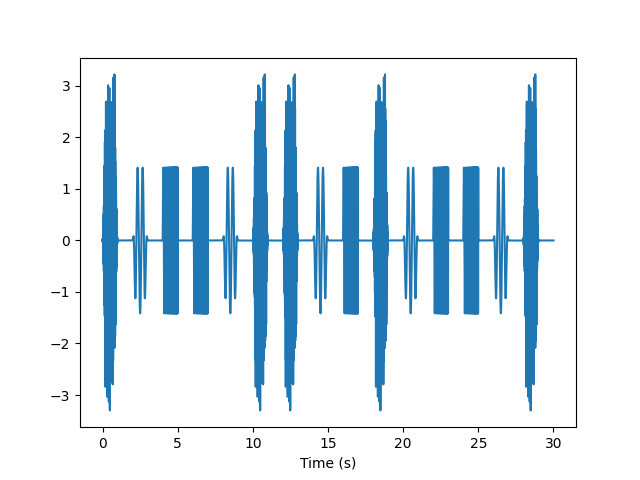
Total running time of the script: (0 minutes 0.395 seconds)